Combining Machine Learning and Molecular Dynamics to Predict P-Glycoprotein Substrates
Por um escritor misterioso
Descrição
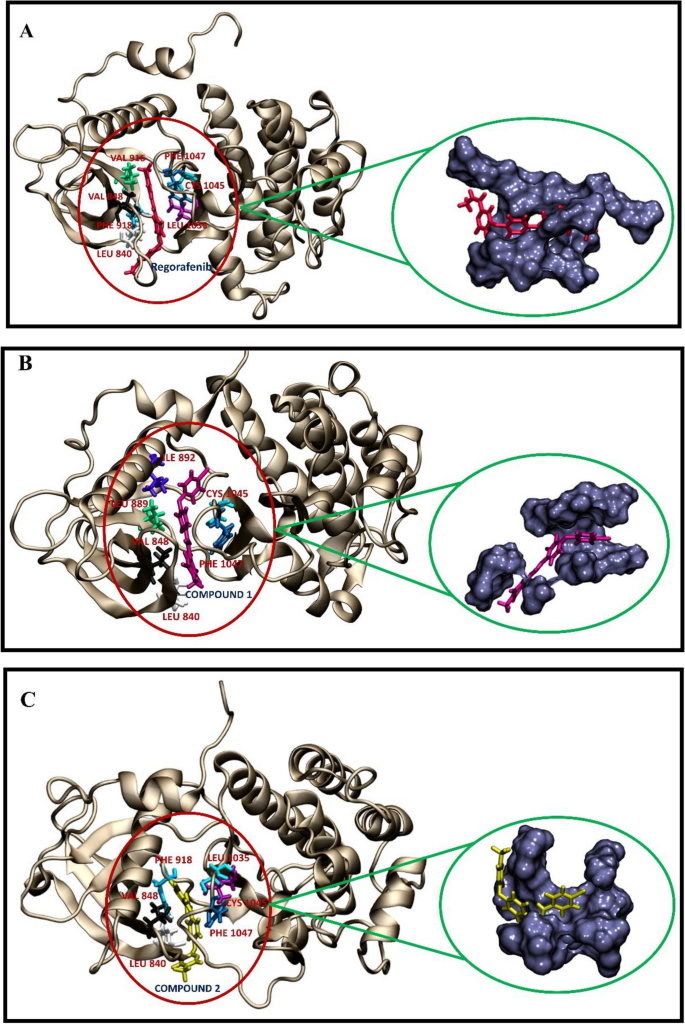
The use of machine learning modeling, virtual screening, molecular docking, and molecular dynamics simulations to identify potential VEGFR2 kinase inhibitors

Interfacial Tension–Temperature–Pressure–Salinity Relationship for the Hydrogen–Brine System under Reservoir Conditions: Integration of Molecular Dynamics and Machine Learning

Multiscale molecular dynamics simulations of lipid interactions with P- glycoprotein in a complex membrane - ScienceDirect
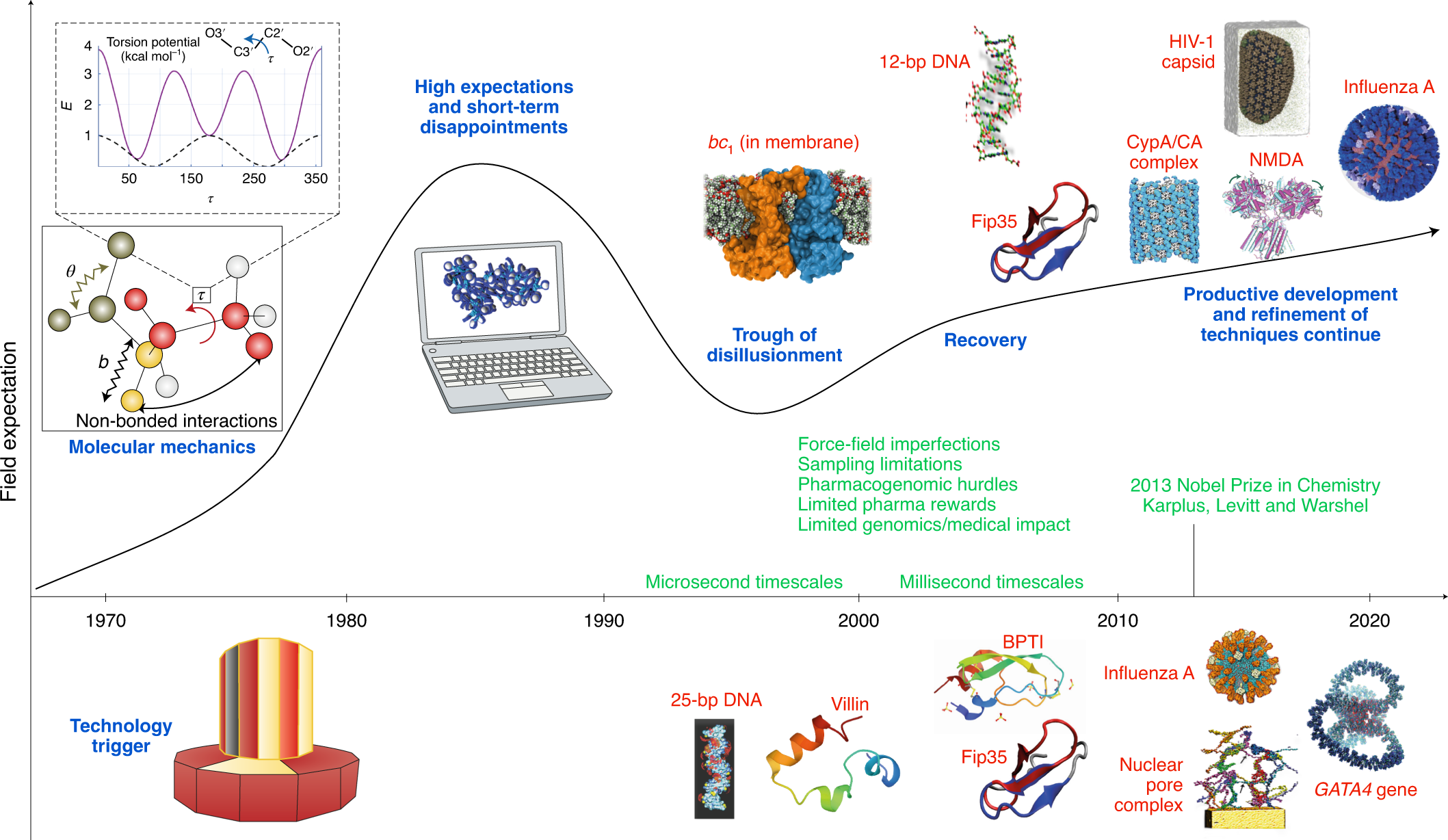
Biomolecular modeling thrives in the age of technology
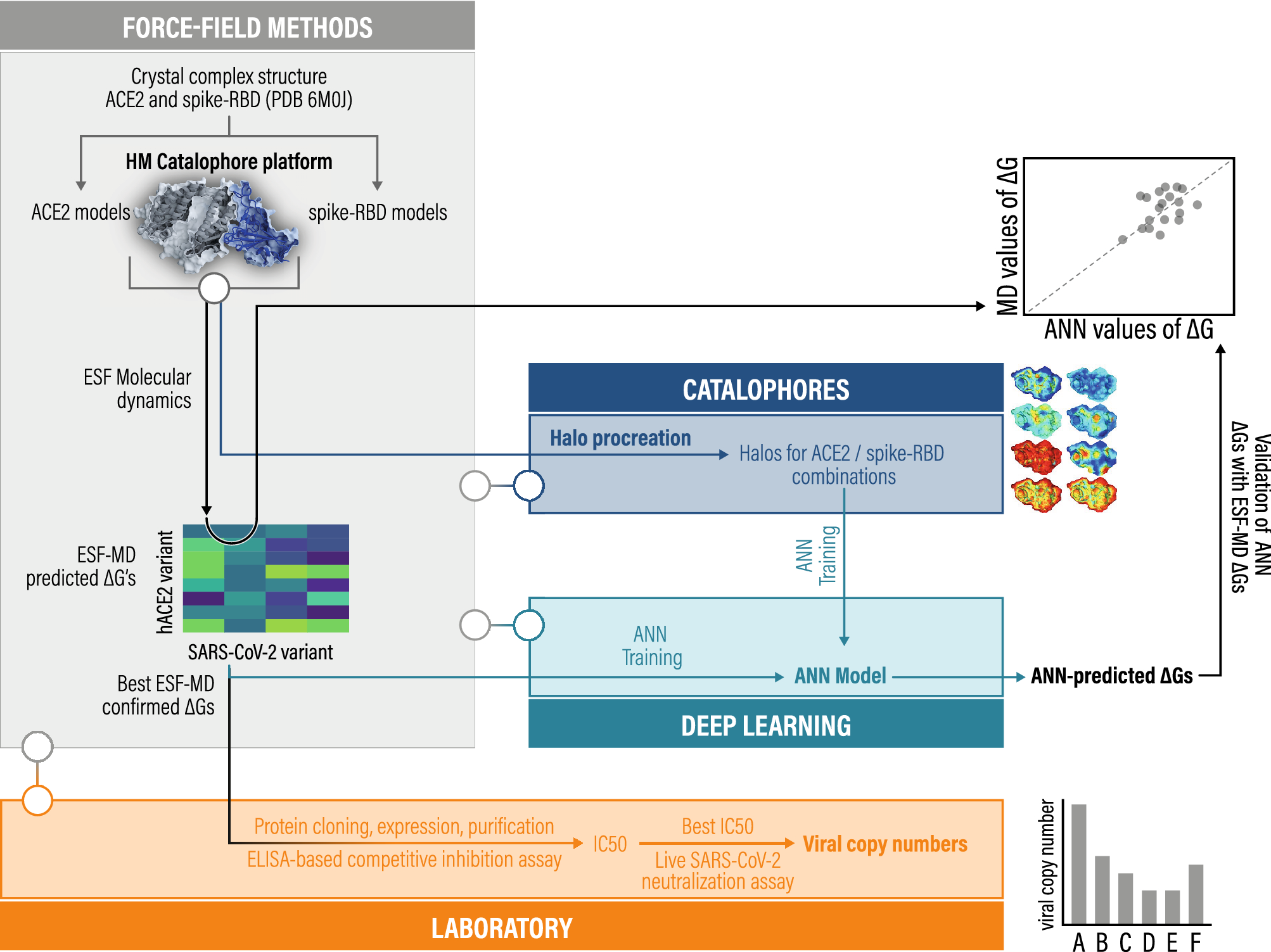
Optimizing variant-specific therapeutic SARS-CoV-2 decoys using deep- learning-guided molecular dynamics simulations

Combining Machine Learning and Molecular Dynamics to Predict P-Glycoprotein Substrates
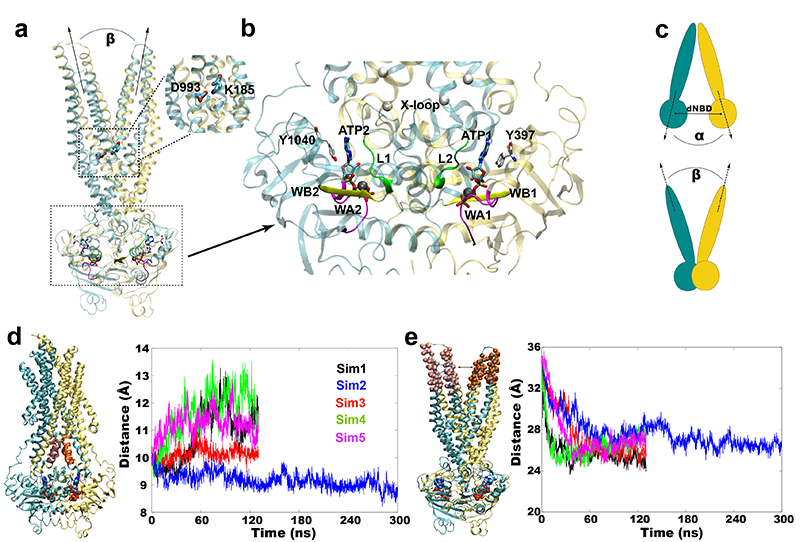
Novel Energy Transduction in P-glycoprotein
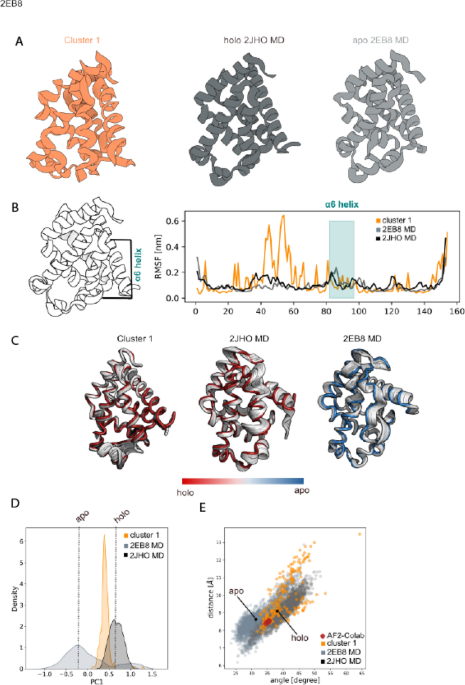
Machine learning/molecular dynamic protein structure prediction approach to investigate the protein conformational ensemble

Deep learning-based molecular dynamics simulation for structure-based drug design against SARS-CoV-2 - ScienceDirect

Combining Machine Learning and Molecular Dynamics to Predict P-Glycoprotein Substrates

Machine learning approaches and their applications in drug discovery and design - Priya - 2022 - Chemical Biology & Drug Design - Wiley Online Library

Combining Machine Learning and Molecular Dynamics to Predict P-Glycoprotein Substrates
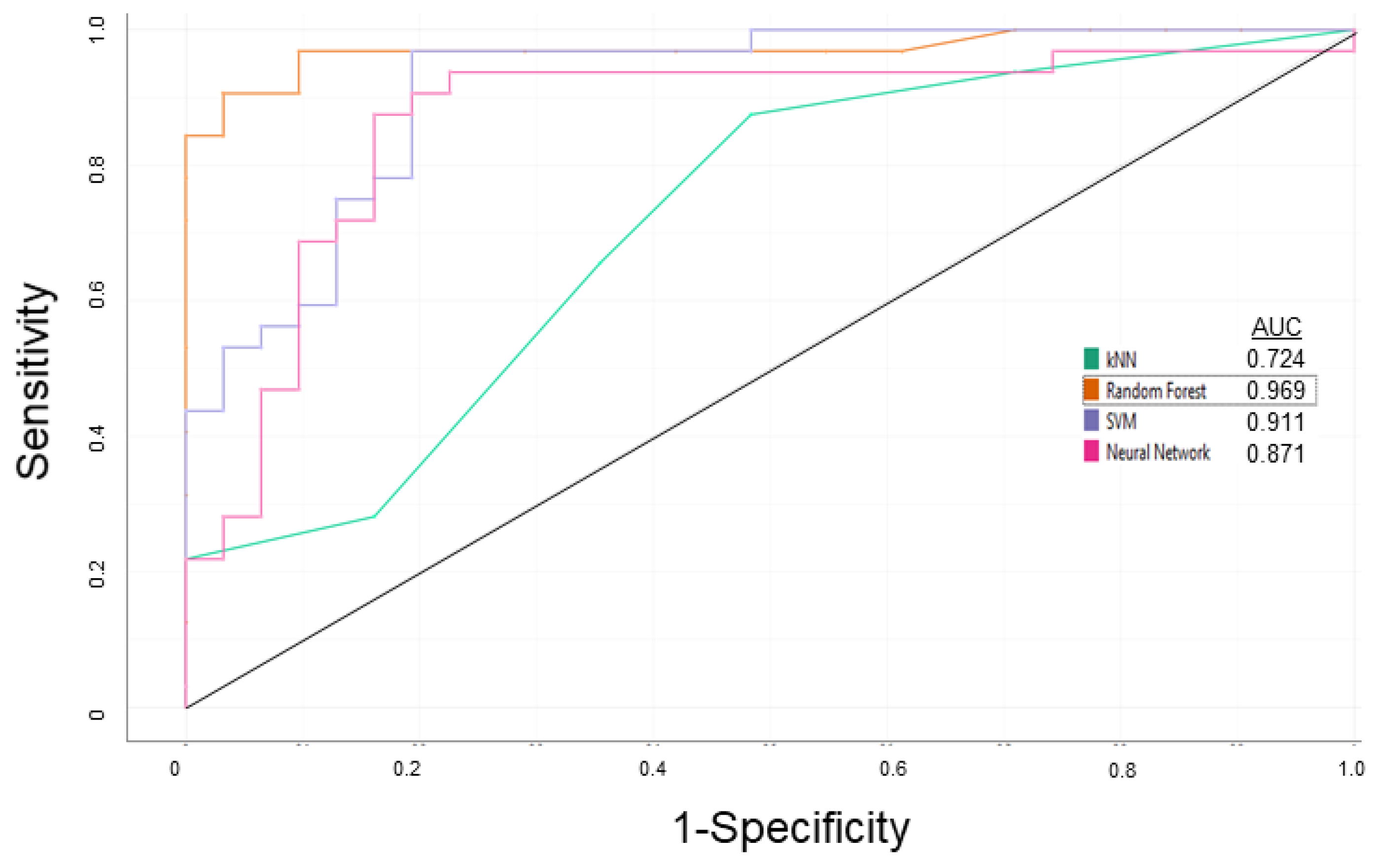
Cells, Free Full-Text
de
por adulto (o preço varia de acordo com o tamanho do grupo)